DICOM files aren’t just for hospitals anymore. These medical imaging files contain rich metadata that developers can use in surprising ways.
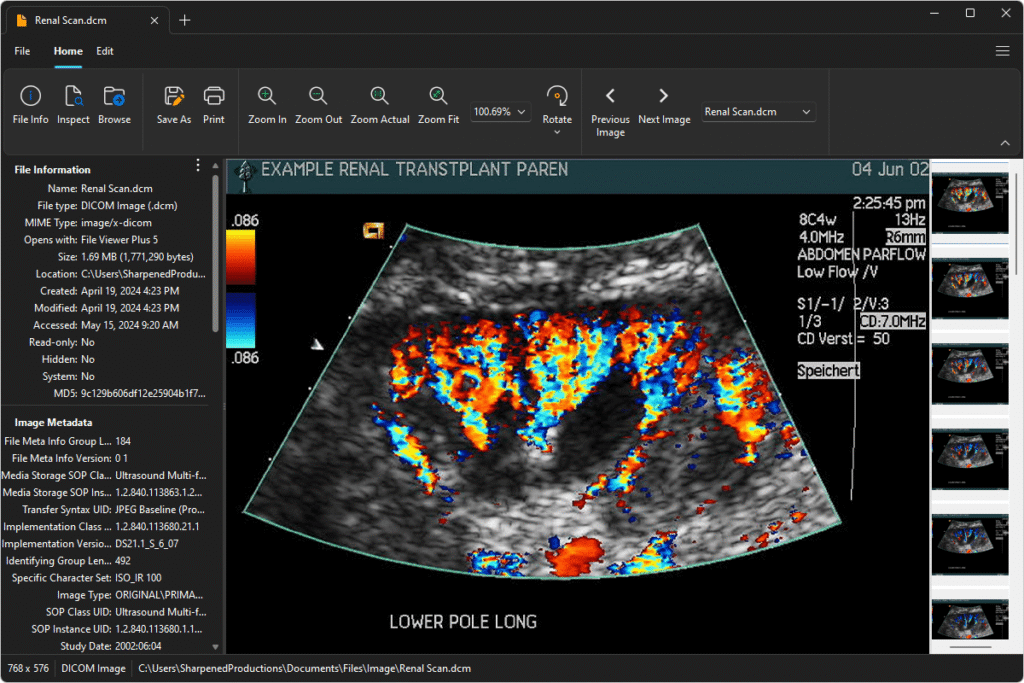
Whether you’re building image analysis tools or how to view DICOM files applications, understanding metadata extraction opens up new possibilities for your projects.
What Makes DICOM Metadata Valuable
DICOM metadata goes far beyond patient information. Each file contains technical details about imaging equipment, acquisition parameters, and processing history.
This data proves useful for developers working on image quality assessment, equipment monitoring, or research applications.
The metadata structure follows specific standards, making it reliable for automated processing.
You’ll find consistent data types and formats across different imaging devices and manufacturers.
Core Components of DICOM Metadata
DICOM files store metadata in a structured format using tags and values. Each tag represents a specific piece of information, from basic details like image dimensions to complex technical parameters.
Key metadata categories include:
- Image Properties: Dimensions, pixel spacing, bit depth
- Acquisition Parameters: Equipment settings, timestamps, protocols
- Processing History: Applied filters, compression methods, calibration data
- Technical Specifications: Device manufacturer, software versions, field strength
Understanding these categories helps you identify which metadata serves your specific application needs.
| Metadata Type | Common Use Cases | Data Format |
| Image Properties | Quality control, processing optimization | Numerical values, strings |
| Acquisition Parameters | Equipment monitoring, protocol analysis | Mixed data types |
| Processing History | Workflow tracking, audit trails | Structured sequences |
Extraction Methods and Tools
Python offers several libraries for DICOM metadata extraction. Pydicom remains the most popular choice due to its comprehensive feature set and active development community.
Here’s how extraction typically works:
import pydicom
# Load DICOM file
ds = pydicom.dcmread(‘sample.dcm’)
# Extract specific metadata
patient_id = ds.PatientID
study_date = ds.StudyDate
image_dimensions = (ds.Rows, ds.Columns)
Alternative approaches include:
- GDCM: C++ library with Python bindings for high-performance processing
- SimpleITK: Medical image analysis toolkit with DICOM support
- DCMTK: Command-line tools for batch processing
Each tool has strengths depending on your project requirements. Pydicom works well for most applications, while GDCM handles large-scale processing better.
Conversion Strategies for Different Formats
Converting DICOM metadata requires careful consideration of target formats. JSON works well for web applications and APIs, while CSV suits data analysis workflows.
JSON conversion preserves complex nested structures:
import json
# Convert to JSON
metadata_dict = {
‘patient_info’: {
‘id’: str(ds.PatientID),
‘age’: str(ds.PatientAge)
},
‘image_specs’: {
‘rows’: int(ds.Rows),
‘columns’: int(ds.Columns)
}
}
json_output = json.dumps(metadata_dict, indent=2)
CSV works better for tabular data analysis. You’ll need to flatten nested structures and handle missing values appropriately.
Real-World Application Examples
Quality Control Systems: Manufacturing companies use DICOM metadata to monitor imaging equipment performance. By tracking acquisition parameters over time, they identify equipment drift before it affects product quality.
Research Data Management: Academic institutions extract metadata to organize large imaging datasets. This enables efficient searching and filtering based on technical parameters rather than just file names.
Image Processing Optimization: Software developers use metadata to automatically adjust processing algorithms. Knowing the original acquisition settings helps optimize enhancement filters and noise reduction techniques.
Equipment Utilization Analysis: Healthcare technology companies analyze metadata to understand how their equipment gets used in different environments. This data informs product development and support strategies.
Common Challenges and Solutions
Missing or inconsistent metadata poses the biggest challenge. Different manufacturers implement DICOM standards differently, leading to variations in available tags.
Build robust error handling into your extraction code:
def safe_extract(ds, tag, default=’Unknown’):
try:
return str(getattr(ds, tag))
except AttributeError:
return default
Large file processing can overwhelm system resources. Consider these optimization strategies:
- Process metadata only (skip pixel data)
- Use streaming readers for large datasets
- Implement batch processing with progress tracking
Data privacy concerns require careful handling of any remaining patient information. Always verify your extraction process removes or anonymizes sensitive data before using it in non-medical applications.
Performance Optimization Tips
Memory management becomes critical when processing many files. Load only the metadata you need and dispose of objects promptly.
Parallel processing speeds up batch operations significantly. Python’s multiprocessing library works well for CPU-bound metadata extraction tasks.
| Processing Method | Files/Second | Memory Usage | Best For |
| Sequential | 10-20 | Low | Small datasets |
| Parallel (4 cores) | 40-80 | Moderate | Medium datasets |
| Streaming | 5-15 | Very Low | Large individual files |
Caching strategies help when you need to access the same metadata repeatedly. Store extracted data in a database or fast file format to avoid reprocessing.
Getting Started with Your Project
Start small with a single DICOM file to understand the metadata structure. Use pydicom’s built-in methods to explore available tags before writing extraction code.
Test your extraction logic with files from different sources. This reveals inconsistencies early in development and helps you build more robust error handling.
Consider your output format carefully. JSON is well-suited for web applications, while CSV is ideal for data analysis workflows. Choose based on how you’ll use the extracted metadata.
Remember that how to view dicom files applications often need both image data and metadata. Plan your extraction process to support both use cases efficiently.
